⇒ Smart pre-processing of force curves – load force curves and access guided tools for denoising (median or Gaussian filter), normalization (base-line correction) and calibration (cantilever and spring constant settings).
⇒ Force curve visualization – display attract and/or retract curves – select axis units: X distance, separation or time; Y deflection (in V or nm) or force in nN.
⇒ Automatic event detection and parameter calculation – detect and display adhesion events automatically (with optional fine tuning of non-zero attraction point and retraction point cursors) – calculate event parameters: distance, force, slope, distance difference, force difference.
⇒ Calculate Young’s modulus, energies, stiffness, etc. and indentation models (Hertz, DMT, JKR, Sneddon etc.)
⇒ Wormlike chain (WLC) models – generate a wormlike chain (WLC) model for protein unfolding and calculate the persistence length, contour length and estimated spring constant at each event.
⇒ Analysis of series of force curves – grid view of all force curves in a series – scroll through a series of force curves and display parameters for each curve in the series – generate statistics (control charts, scatter plots, histograms) on any parameter.
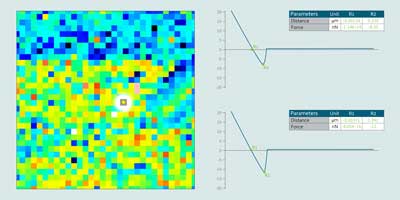
⇒ Visualization of force volume data – display an interactive parameter map representing the value of a parameter calculated on each element of a force volume dataset.
⇒ Filter force volume and extract – smooth force volume data by removing noise and extract slices or individual force curves interactively.
Included in:
- MountainsSPIP® Premium
- MountainsLab® Premium
Available with:
- MountainsSPIP® Starter
- MountainsSPIP® Expert
- MountainsSpectral® Expert
- MountainsSpectral® Premium
- MountainsLab® Expert